
I lead a translational genomics program with the goals of 1) linking cancer genome alterations to clinical outcome; 2) monitoring shifts in cancer and immune cell populations during treatment; and 3) enabling comprehensive genomics as a routine medical test. To achieve these goals, we profile tumours and blood using cell-free DNA, immune repertoire, single-cell and whole-genome sequencing. We also develop software to share these data around the world. Our work advances the knowledgebase and infrastructure necessary to deliver genomic medicine to cancer patients worldwide.
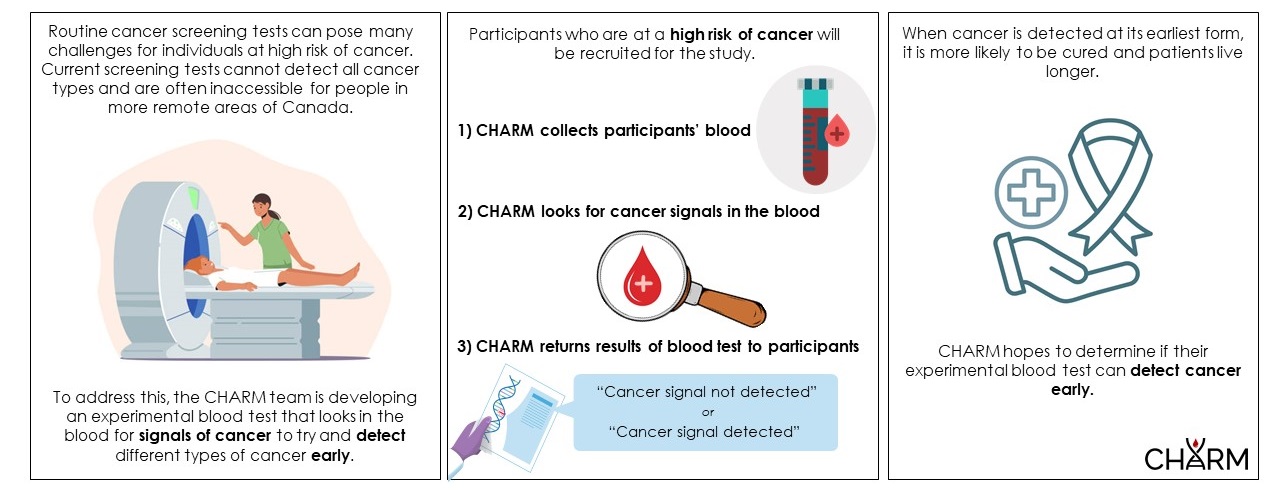
Cell-free DNA in Hereditary And High- Risk Malignancies (CHARM)2: Evaluating the Performance of a cfDNA Blood Test for Early Cancer Detection
Study Status: active
Institute: Princess Margaret Cancer Centre
Health Conditions:
Cancer
External Links:
Dr. Trevor Pugh's research program is focused on understanding clinical implications of clonal shifts in cancer and non-cancerous cell populations during treatment, most recently using cell-free DNA, immune repertoire, and single cell RNA-seq sequencing. He has contributed to multiple large-scale genomics and data-sharing programs including AACR GENIE and the Terry Fox Canadian Comprehensive Cancer Centre Network. Previously, as a postdoctoral fellow with Dr. Matthew Meyerson at the Dana-Farber Cancer Institute and Broad Institute of Harvard and MIT, Dr. Pugh led landmark cancer genome studies describing three pediatric solid tumours: medulloblastoma, neuroblastoma, and pleuropulmonary blastoma. During this time, he also completed a clinical laboratory fellowship in the Harvard Medical School Genetics Training Program with Dr. Heidi Rehm. Originally from Vancouver, Dr. Pugh received a PhD in Medical Genetics from the University of British Columbia.
- Lead, Clinical Genomics Program, Princess Margaret Cancer Centre
- Professor, Department of Medical Biophysics, University of Toronto
- Senior Investigator and Director, Genomics, Ontario Institute for Cancer Research