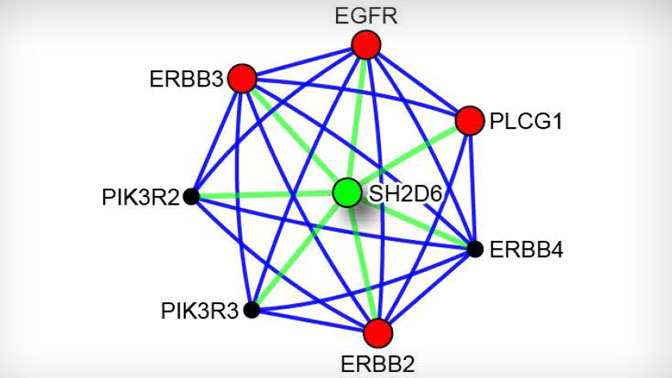
Imagine going to a museum where none of the artifacts are labelled. Without any knowledge of the context in which the artifacts were used or where they came from, an exhibit would look like a collection of random objects.
In the past several decades, researchers have amassed large collections of information describing the components of cells and how they work together. However, much of the context has been missing from this information, which has prevented researchers from fully understanding the role of a cell’s components in health and disease.
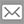
To help address this, Dr. Igor Jurisica, Senior Scientist at Krembil Research Institute, has been ‘putting labels’ on the interactions between proteins, the cellular machinery that performs much of the essential functions of the cell. In 2005, he created a database of protein interactions that has evolved and expanded over the years. In 2015, it included proteins from humans and five other species, as well as the tissues in which specific interactions occur.
His team recently reported a substantial expansion of their database in the journal Nucleic Acids Research. The new features include protein interactions for 12 more species to support biomedical, veterinary and agricultural research, and three new types of context information: where these protein interactions occur in the cell, the diseases they are associated with, and whether they might respond to a potential drug.
“The Integrated Interactions Database, or IID, is one of the broadest and largest physical interaction databases,” explains Dr. Jurisica. “We’ve also included features that enable researchers to map the networks of protein interactions for each species, identify the key proteins in these networks and determine the conditions under which an interaction network is physiologically important.”
With this additional information, the 4.8 million protein-protein interactions in the database will help researchers better understand the molecular mechanisms behind diseases and treatments, and develop new drugs faster.
This work was supported by the Krembil Foundation, Ontario Research Fund, Natural Sciences and Engineering Research Council of Canada, Canada Foundation for Innovation, Canada Research Chairs Program, Toronto General & Western Hospital Foundation and IBM.
Kotlyar M, Pastrello C, Malik Z, Jurisica I. IID 2018 update: context-specific physical protein-protein interactions in human, model organisms and domesticated species. Nucleic Acids Res. 2018 Nov 8. doi: 10.1093/nar/gky1037.

Dr. Igor Jurisica, Senior Scientist at Krembil Research Institute.