Researchers at UHN's McEwen Stem Cell Institute shed light on the role of cardiac fibroblasts in heart development and disease, key cells for improving treatments for heart-related conditions, specifically those affecting the heart muscle.
Cardiac fibroblasts make up approximately 15–30% of all heart cells and are essential for both fetal heart development and maintenance of the adult heart structure. The majority of the fibroblast population is derived from the outer layer of the heart, called the epicardium.
"There's a lack of laboratory models that accurately replicate the formation and function of the epicardium, which hinders our understanding of heart development, disease mechanisms, and the advancement of new therapies," explains Dr. Gordon Keller, Director of the McEwen Stem Cell Institute and senior author of this study.
To address this gap, the team developed cardiac organoids, three-dimensional pluripotent stem cell-derived structures, designed to mimic key aspects of human heart development and function. These organoids spontaneously form the functional muscle tissue and epicardium found in the heart, providing a platform to study cardiac cell interactions in a more physiologically relevant environment.
Using advanced techniques to monitor cell differentiation and maturation, researchers discovered that interactions between epicardial cells and heart muscle cells (cardiomyocytes) within the organoids resulted in the specification and migration of fibroblasts, in a pattern similar to that observed in developing human hearts.
"Our study demonstrates that this organoid platform serves as an advanced model system for studying multicellular mechanisms underlying heart disease," explains Dr. Ian Fernandes Scientific Associate at McEwen Stem Cell Institute and first author of this study.
Moreover, single-cell RNA sequencing, a cutting-edge technique that allows scientists to study the molecular profile of individual cells, revealed significant diversity within the populations of fibroblasts and cardiomyocytes in the cardiac organoids. This diversity resembled what is seen in both healthy and diseased adult hearts.
Researchers also identified distinct subpopulations of fibroblasts, including a group expressing the CD9 protein, which has been associated with reparative functions.
"The identification of distinct subpopulations of cells within the heart organoids, especially those with reparative properties like the CD9+ fibroblasts, opens up new avenues for understanding and potentially treating heart diseases," adds Dr. Keller. "By targeting these specific cell types, we may develop more precise and effective therapies for various heart-related conditions."
This research opens avenues for understanding cardiac development, disease mechanisms, and potential regenerative therapies, promising advancements in heart-related treatments. Additionally, the ability to isolate and co-transplant specific fibroblast subpopulations with cardiomyocytes in damaged hearts could hold potential for generating new heart muscle and improving heart function.
This work was supported by the Canadian Institutes of Health Research and UHN Foundation. Dr. Gordon Keller is a Professor at the Department of Medical Biophysics at the University of Toronto.
G.M.K. is a scientific co-founder and paid consultant for BlueRock Therapeutics LP and a paid consultant for VistaGen Therapeutics.
Fernandes, I., Funakoshi, S., Hamidzada, H. et al. Modeling cardiac fibroblast heterogeneity from human pluripotent stem cell-derived epicardial cells. Nat Commun 14, 8183 (2023). https://doi.org/10.1038/s41467-023-43312-0
Researchers from UHN’s KITE Research Institute offer new hope for epilepsy research as they develop deep-learning models to predict epileptic seizures.
Epilepsy, one of the world’s most prevalent neurological disorders, affects over 50 million people worldwide. Characterized by the sudden onset of seizures, epilepsy can lead to serious physical injury and even death. The ability to predict the onset of epileptic seizures can significantly reduce injury and improve quality of life.
A team led by Dr. Shehroz Khan, KITE Scientist and senior author of the study, focused on leveraging deep learning models to analyze electroencephalogram (EEG) data. EEG is a test that uses small electrodes to measure brain activity and serves as a vital tool for understanding seizure onset.
“Deep learning models are advanced computer algorithms that learn to recognize patterns and make predictions by processing large amounts of complex data. By using these models to distinguish pre-seizure EEG patterns, we can help epilepsy patients and their caregivers anticipate seizures and take preventive measures,” states Dr. Khan.
Using a combination of supervised and unsupervised deep learning approaches, the researchers trained the learning models to identify subtle changes in brain activity preceding seizures.
“Supervised deep learning involves using labelled data where seizure occurrence is known. On the other hand, unsupervised deep learning allows the model to learn predictive patterns from unlabeled data on its own,” explains Zakary Georgis-Yap, a previous master's student in Dr. Khan’s lab and first author of the study. “The advantage of unsupervised learning models is that they do not require comprehensively labelled data—which can be challenging and time-consuming to obtain.”
To evaluate the effectiveness of their models, the researchers conducted extensive testing on two large seizure datasets containing EEG-recorded data from 40 patients.
The results of the study were promising, showcasing the feasibility of both supervised and unsupervised approaches in seizure prediction. However, prediction results for both models varied across datasets, patients, and learning approaches, highlighting the considerable variability in pre-seizure brain activity between individuals.
“While there is still work to be done, our research represents a significant step forward in the field of epilepsy management,” concludes Dr. Khan. “By harnessing the potential of deep learning, we have the opportunity to develop personalized therapeutic interventions and ultimately save lives.”
This work was supported by the Natural Sciences and Engineering Research Council of Canada, the Data Science Institute at the University of Toronto and UHN Foundation. Dr. Shehroz Khan is an Assistant Professor at the University of Toronto’s Institute of Biomedical Engineering.
#Georgis-Yap, Z., Popovic, M.R. & #Khan, S.S. Supervised and Unsupervised Deep Learning Approaches for EEG Seizure Prediction. J Healthc Inform Res. 2024 Jan 4. DOI: 10.1007/s41666-024-00160-x
#Zakary Georgis-Yap and Dr. Shehroz Khan contributed equally to the study.
Researchers from Princess Margaret Cancer Centre (PM) have developed an approach to predict outcomes in patients treated with pembrolizumab—an immunotherapy drug used to treat various cancers such as head and neck, breast, and skin cancer.
Analysis for this study was performed using blood samples collected from patients with different cancer types treated with pembrolizumab in the INSPIRE trial, an investigator-initiated study that was designed and carried out at PM.
Pembrolizumab is a type of antibody that attaches to a protein known as programmed death 1 (PD-1), hindering its activity. PD-1 acts as a checkpoint for the immune system, preventing it from becoming overactive and attacking healthy cells. But in the case of cancer, PD-1 can also act as a brake to prevent the immune system from attacking cancer cells. Therefore, inhibiting PD-1 releases the brakes on the immune system and allows it to recognize and attack cancer cells more effectively.
“Although pembrolizumab is used to target many cancers, some patients may develop resistance to the drug,” says Dr. Stutheit-Zhao, Postdoctoral Researcher at PM and co-first author of the study. “Biomarkers like circulating tumour DNA (ctDNA) can provide insights into the effectiveness of treatments without the need for invasive procedures. However, many current ctDNA analyses are so-called tumour-informed, thus requiring molecular sequencing of patients' tumour tissue to identify alterations that can be tracked in the blood.”
ctDNA are pieces of tumour DNA that circulate in the bloodstream. Previous studies—including a clinical trial from this team—have shown that changes in levels of ctDNA using a tumour-informed assay early on during the treatment course can predict pembrolizumab treatment success and patient survival. However, as this assay also requires tumour samples, it may not be feasible for some patients as tumours may be in challenging locations to biopsy, or biopsy specimens may contain insufficient tumour cells for genetic analysis.
“We wanted to investigate a way to predict treatment outcome using ctDNA alone without needing to analyze a tumour biopsy,” says Dr. Enrique Sanz-Garcia, Clinician Investigator at PM and co-first author of the study. “DNA from tumours is known to be enriched for a type of modification, called methylation. Therefore, we analyzed methylated cell-free DNA from 204 plasma samples from 87 patients before and during treatment with pembrolizumab from the INSPIRE trial. This particular assay does not require access to patients’ tumour samples.”
Previous studies have demonstrated the ability of a technique, first introduced by scientists at UHN, that isolates and analyzes this methylated DNA, called cell-free methylated DNA immunoprecipitation and sequencing (cfMeDIP-seq), to estimate ctDNA levels. This technique can also quantify DNA fragment lengths, which are known to be shorter in tumour tissue compared to DNA derived from normal tissue.
The team therefore retrospectively tested whether methylation and fragmentation status of cell-free DNA using cfMeDIP-seq analysis can monitor the response to pembrolizumab in the INSPIRE study.
“Our findings indicated a strong correlation between the ctDNA analysis of methylation patterns and DNA fragmentation in blood samples alone with tumour-informed ctDNA analysis,” says Dr. Lilllian Siu, Senior Scientist at PM and senior author of the study. “Importantly, methylation analysis predicted overall survival and progression-free survival, and fragment length analysis was able to predict overall survival in patients treated with pembrolizumab.”
Using statistical analysis, the team found that a decrease in cancer-specific methylation over the first six weeks of pembrolizumab treatment was associated with a 60% lower risk of death and a 65% lower risk of disease progression compared to the reference group. For fragmentation analysis, a decrease in fragment length score over time was significantly associated with a 60% lower risk of death.
This suggests that analysis of circulating DNA assayed by cfMeDIP-seq yields promising tumour-naïve (i.e. not requiring access to tumour samples) predictive biomarkers for response to pembrolizumab.
“This is the first reported examination of methylated cell-free DNA in advanced cancer patients during treatment with pembrolizumab and one of the first direct comparisons between ctDNA analysis with and without tumour samples,” adds Dr. Trevor Pugh, Senior Scientist at PM and co-senior author of the study.
This study, published in Cancer Discovery, the lead journal of the American Association for Cancer Research, reveals the potential of a minimally invasive blood test (i.e., a liquid biopsy) for predicting response to cancer treatments. This could enable earlier response assessment by doctors and lead to prompt redirection to next-line treatment options in non-responders before radiological assessments.
Circulating tumour DNA (ctDNA) can be analyzed using blood tests as part of a liquid biopsy. Liquid biopsies are blood tests used to detect cancer cells or other markers of disease.
This work was supported by The Princess Margaret Cancer Foundation, the Ontario Institute for Cancer Research, the Terry Fox Research Institute, the Princess Margaret Cancer Centre Global Oncology Program, and the Cancer Research Institute.
Dr. Enrique Sanz-Garcia is an Assistant Professor at the University of Toronto (U of T). Dr. Lilian Siu is a Professor of Medicine at U of T and holds the BMO Chair in Precision Cancer Genomics. Dr. Pugh is a Professor in the Department of Medical Biophysics at U of T and is a tier 2 Canada Research Chair in Biological Physics.
Dr. Sanz Garcia reported research funding from Novartis. Dr. Siu reported either a leadership role, financial interest in, or a consulting role with Treadwell Therapeutics, Agios, Merck, AstraZeneca/MedImmune, Roche, Voronoi Health Analytics, Oncorus, GlaxoSmithKline, Seattle Genetics, Arvinas, Navire, Janpix, Relay Therapeutics, Daiichi Sankyo/UCB Japan, Janssen, Hookipa Pharma, InterRNA, Tessa Therapeutics, Sanofi, Amgen, and research funding from Bristol Myers Squibb (Inst),Genentech/Roche (Inst), GlaxoSmithKline (Inst), Merck (Inst), Novartis (Inst), Pfizer (Inst), AstraZeneca (Inst), Boehringer Ingelheim (Inst), Bayer (Inst), Amgen (Inst), Astellas Pharma (Inst), Shattuck Labs (Inst), Symphogen (Inst), AVID Radiopharmaceuticals (Inst), Mirati Therapeutics (Inst), Intensity Therapeutics (Inst), Karyopharm Therapeutics (Inst). Dr. Pugh has provided consultation for AstraZeneca, Chrysalis Biomedical Advisors, Merck, and SAGA Diagnostics (compensated); and receives research support (institutional) from Roche/Genentech.
For a full list of competing interests, see manuscript.
Stutheit-Zhao EY*, Sanz-Garcia E*, Liu ZA, Wong D, Marsh K, Abdul Razak AR, Spreafico A, Bedard PL, Hansen AR, Lheureux S, Torti D, Lam B, Yang SYC, Burgener J, Luo P, Zeng Y, Cheng N, Awadalla P, Bratman SV, Ohashi PS, Pugh TJ, Siu LL. Early changes in tumor-naive cell-free methylomes and fragmentomes predict outcomes in pembrolizumab-treated solid tumors. Cancer Discov. 2024 Feb 23. doi: 10.1158/2159-8290.CD-23-1060. Epub ahead of print. PMID: 38393391.
*authors contributed equally to this article.
Researchers from UHN’s KITE Research Institute have investigated a new affordable and clinically accessible training system for improving the standing balance of spinal cord injury patients.
Spinal cord injury affects approximately 85,000 Canadians every year. Individuals with incomplete spinal cord injury can regain their ability to walk; however, a majority of these individuals experience falls at least once a year. Falling can reduce mobility, physical activity, and also quality of life.
Balance therapy helps increase muscle strength and reaction but relies on an individual’s vision to help maintain standing balance. Visual feedback balance training (VFBT) is a training method that incorporates visual cues to help improve balance and postural control. It involves shifting your body toward a target location on a screen.
Functional electrical stimulation is an additional rehabilitation technique that uses electrical impulses to activate muscles, helping individuals regain movement and improve function. Researchers have explored combining VFBT with functional electrical stimulation as a comprehensive rehabilitation approach for improving balance control. However, these rehabilitation systems traditionally rely on costly force plates to measure a participant’s movement.
The team led by Dr. Kei Masani, KITE Senior Scientist and senior author of the study, investigated the integration of low-cost and portable sensors like a depth camera and pressure mat, which use motion tracking and distribution of pressure, respectively, to analyze movement.
The effectiveness of these sensors was put to the test by measuring the movements of ten able-bodied participants, with no history of neurological disorders, as they completed balance rehabilitation exercises using the combined VFBT and functional electrical stimulation system. Researchers found that the depth camera outperformed the pressure mat, showing higher accuracy and lower error relative to the force plate, in capturing crucial balance and movement measures.
These results suggest that the depth camera could replace the force plate in the VFBT and functional electrical stimulation rehabilitation system.
Derrick Lim, PhD candidate in the lab of Dr. Masani and first author of the study, is enthusiastic about these findings stating, “This study marks a significant step to making rehabilitation more accessible and effective for individuals with spinal cord injuries. The use of affordable sensors opens doors to broader implementation in clinical settings, ultimately improving the quality of life for patients.”
Future work will focus on using this system with individuals who have experienced spinal cord injury and have poorer balance capabilities. It has potential applications in other populations who experience neurological damage resulting in balance impairments, such as adults living with stroke.
This work was supported through the Collaborative Health Research Projects program, a joint initiative between both, the Canadian Institutes of Health Research (CIHR) and the Natural Sciences and Engineering Research Council of Canada (NSERC), as well as UHN Foundation. Dr. Kei Masani is an Associate Professor at the Institute of Biomedical Engineering at the University of Toronto.
Lim D, Pei W, Lee JW, Musselman KE, Masani K. Feasibility of using a depth camera or pressure mat for visual feedback balance training with functional electrical stimulation. Biomed Eng Online. 2024 Feb 12. doi: 10.1186/s12938-023-01191-y.
Research from UHN’s Krembil Brain Institute has revealed potential disease-modifying effects of deep brain stimulation in Parkinson’s disease (PD), shedding new light on an old treatment.
PD is a neurodegenerative disease that affects millions of people worldwide. The disease is characterized by motor symptoms, such as tremors, stiffness, slowed movement, and impaired balance. These hallmark symptoms are caused by a progressive loss of neurons in an area of the brain that controls movement, called the basal ganglia.
For some people with PD, severe motor symptoms can be treated with a surgical procedure known as deep brain stimulation (DBS). This therapy involves implanting electrodes into specific brain areas where they deliver electrical impulses that correct abnormal neuron activity.
Despite the known benefits of DBS for relieving PD symptoms, there are still many unknowns surrounding how the therapy works and whether it can alter the course of the disease.
In a recent study published in Brain Stimulation, Krembil Brain Institute researchers Drs. Suneil and Lorraine Kalia provided evidence that DBS may serve as more than just a symptomatic treatment—it might actually alter underlying disease processes.
“A driving factor in most neurodegenerative diseases is the accumulation of misfolded proteins within or outside of brain cells,” explains Dr. Suneil Kalia, a Krembil Senior Scientist and co-lead of the study. “In PD, we see a buildup of a misfolded form of the protein alpha-synuclein (α-Syn), which disrupts neuron communication and eventually causes neuron death.”
According to Dr. Suneil Kalia, the search is on for treatments that can disrupt α-Syn production and aggregation, thereby slowing or even preventing PD progression.
To determine whether DBS has such disease-modifying effects, the Kalia labs examined whether high-frequency electrical stimulation, akin to that delivered by DBS electrodes, alters α-Syn.
“When we stimulated cultured brain cells, we saw significantly less expression and buildup of aggregated α-Syn. Within the brain, when we stimulated a small region of the basal ganglia called the substantia nigra pars compacta, we saw a lower overall level of α-Syn and less downstream effects,” says Dr. Lorraine Kalia, a Krembil Senior Scientist, Neurologist and co-lead of the study.
These findings hint at the potential of DBS to protect neurons and slow down the disease.
Interestingly, the team saw this benefit only when they simulated the substantia nigra pars compacta and not a nearby region that clinicians more commonly target in DBS.
Dr. Suneil Kalia is also a Neurosurgeon at Krembil Brain Institute and, together with his two colleagues, performs the largest number of DBS surgeries in Canada. “We regularly apply DBS to one of three functionally connected regions of the basal ganglia, depending on the patient and their specific symptoms—most commonly the subthalamic nucleus. DBS in this region can dramatically improve a patient’s motor symptoms, so we were somewhat surprised to see that it had no effect on α-Syn levels,” explains Dr. Suneil Kalia.
“This finding tells us that the disease-modifying actions of DBS may depend to some extent on the brain region being targeted—this will be an important consideration when optimizing treatment plans for individual patients,” he adds.
Dr. Eun Jung Lee and Dr. David Aguirre-Padilla are co-first authors of the study and former Postdoctoral Researchers in the Kalia labs. “DBS has been used to treat PD since the late 1990s but it is only now that we are learning about its disease-modifying properties. Our findings underscore the potential of harnessing existing therapies for neurological diseases in new ways,” explains Dr. Aguirre-Padilla.
Although more research is needed to determine exactly how DBS alters α-Syn, this study offers hope for millions of patients and their caregivers.
“The better we understand how DBS works, the more we can refine the therapy to enhance its benefits for each patient. This approach could really change the landscape of PD treatment,” concludes Dr. Lee.
PD affects more than 100,000 Canadians. DBS is a valuable treatment option for a subset of patients whose motor symptoms cannot be controlled by medication alone.
This work was supported by Parkinson Canada, Krembil Research Institute and UHN Foundation. Dr. Suneil Kalia is an Associate Professor of Surgery and holds the R. R. Tasker Chair in Stereotactic and Functional Neurosurgery at UHN. Dr. Lorraine Kalia is an Associate Professor of Medicine and holds the Wolfond-Krembil Chair in Parkinson’s Disease Research at UHN.
Lee EJ, Aguirre-Padilla DH, Fomenko A, Pawar G, Kapadia M, George J, Lozano AM, Hamani C, Kalia LV, Kalia SK. Reduction of alpha-synuclein oligomers in preclinical models of Parkinson’s disease by electrical stimulation in vitro and deep brain stimulation in vivo. Brain Stimul. 2024 Feb 10. Doi: 10.1016/j.brs.2024.02.005.
I am currently a postdoctoral researcher at UHN’s KITE Research Institute (KITE). My two research projects focus on upper limb rehabilitation using novel robotic training and neuromodulation modalities after stroke and spinal cord injury.
I am also a member of the UHN Research IDEA Curriculum Advisory Sub-Committee. As a woman from a minority background, this has been a great opportunity for me to share my experience working in diverse healthcare systems. I am particularly interested in helping trainees integrate IDEA principles into their work and understand how to navigate working in diverse clinical and research environments. Given the diversity of the Canadian population, researchers need to understand how to effectively address gaps in health care and meet the needs of our patients.
I have been at UHN for 3 years. I am originally from the Republic of Mauritius where I studied to become a physiotherapist. I have an MSc in Physiotherapy-Neurorehabilitation from the University of Nottingham, United Kingdom, and a PhD in Physiotherapy from the University of Newcastle, Australia. I previously was also faculty at the University of Mauritius for 7 years leading the physiotherapy education program. In 2021, I immigrated to Canada to fulfill my research career aspirations and challenge myself to do a postdoctoral fellowship.
As a postdoctoral researcher at UHN, I am passionate about developing new neurorehabilitation technologies for people with stroke and spinal cord injury. I enjoy the opportunity to work with advanced robotics and neuromodulation modalities that could be pivotal in optimizing recovery for neurological patients. I also enjoy engaging in stimulating scientific discussions with other researchers and trainees. As a clinician and researcher, I find that pursuing a career in health research can be quite challenging; however, working with a great team helps maintain resilience.
I believe that health research is the driving force behind optimal care for our patients and should be prioritized. As health research goes hand-in-hand with healthcare delivery, more effort should be concentrated on overcoming obstacles in translating research into clinical practice.
My mission is to develop rehabilitation approaches to help restore movement for people with neurological conditions and improve their quality of life. My research program specifically focuses on developing complex neurorehabilitation approaches using clinical restorative therapy and advanced rehabilitation technology like robotic training, haptic training using pressure sensors, and neuromodulation.
My research addresses a unique area of research designed to advance our understanding of the complex interactions between sensory function, motor function, and cognitive processes necessary for upper limb function. I am particularly interested in studying young and older stroke survivors to better understand the effects of age on upper limb recovery after stroke. I am also interested in studying people who have reintegrated into the community to understand how we can meet their rehabilitation needs in the long-term.
I feel privileged to conduct my post-doctoral research at KITE and the Toronto Rehabilitation Institute (Toronto Rehab), one of the world’s top-ranked rehabilitation research institutes. I have developed an extensive network of multidisciplinary collaborators to advance my research program beyond traditional rehabilitation to integrate several aspects of biomedical engineering such as artificial intelligence and advanced robotics. I have also met many scientists who have mentored me in various academic pursuits.
Additionally, Toronto Rehab has connections with several research platforms which allows for unique opportunities to connect with leading researchers from across Canada. There is a lot of support provided within the UHN network that makes UHN an excellent environment for researchers to advance health research.
I have enjoyed learning Kathak, which is an Indian form of classical dance, for 9 years and have a degree in Kathak dance. I occasionally practice Kathak whenever I feel inspired. But my main hobby is reading books or listening to podcasts about self-development and personal growth. I also enjoy travelling both locally and abroad, exploring nature like trails and snorkeling.
I think the future of health research lies in integrating advanced technologies such as artificial intelligence, big data, and innovative technological devices. This will help us provide more personalized medicine and rehabilitation for our patients.
The future of health research is even more promising with increasing contributions from diverse researchers, trainees, healthcare leaders, and patient partners. As an immigrant, it is an exciting opportunity for me to be part of this process in Canada. I am excited about collaborating with local and international collaborators to advance the field of physiotherapy and neurorehabilitation. I am also enthusiastic about trialing new rehabilitation technologies that have the potential to revolutionize rehabilitation and impact the lives of patients.
You @TeamUHN is a campaign to highlight the important scientific contributions that research lab staff, trainees and learners, administrative staff, core facilities staff, Research Solutions & Services staff, and volunteers make towards A Healthier World through discovery and innovation. If you’re interested in sharing your story, we invite you to complete this form here (open to UHN staff, trainees and volunteers).
“It is a scary topic when someone tells you that your job is going to be replaced by AI,” says Dr. Phedias Diamandis, a neuropathologist at UHN.
Modern AI's image analysis mirrors a pathologist's clinical skills in examining microscopic tissue images to diagnose diseases. Concerns about AI overtaking pathology have existed for years.
To transform the challenge into opportunities, Phedias stepped foot in the world of AI in 2017, when the concerns began to spread.
“I wanted to see if AI can interpret pathology images like humans are trained to do.”
Drawing upon his neuroscience background and extensive self-learning, Phedias discovered that AI learns in a manner comparable to humans.
"Our perception relies on neural networks in our brain. In the visual system, primary visual centers detect basic shapes like circles and squares, while higher-level centres integrate them into complex objects. Similarly, AI analyzes images by identifying basic shapes through spatial coordinates and checking for spatial distribution to recognize familiar patterns."
With an understanding of how AI works and a goal to apply AI in pathology, Phedias’ team started a research project to train AI to recognize tumour histology slides.
The team fed AI nearly 1 million images collected from over 1,000 brain tumors, each annotated by pathologists. Using deep neural networks, they developed a tool to analyze cell patterns and generate a map highlighting distinct regions of the tumor with their unique histomorphological features.
They call this AI-driven tool “HAVOC” (Histomic Atlases of Variation Of Cancers). HAVOC aims to help researchers better understand tumour heterogeneity–a phenomenon where different regions of a tumour can have different biology, which can lead to different treatment responses and resistance. Understanding cancer variations helps guide personalized treatment and improve precision medicine.
But how well does it work?
The team used whole slide images from six high-grade glioblastomas to test HAVOC’s accuracy. As the images are processed through the deep neural networks, HAVOC extracts the morphology features from small patches of each image and clusters the patches by similarity to mark different regions in the original image.
It turned out that the results obtained by HAVOC are consistent with the interpretations made by human experts and are correlated to the molecular variations in the sample. In some cases, the variations in molecular features predicted by HAVOC could support the use of personalized combination therapies for patients (published in Science Advances).
“HAVOC can assist pathologists to understand tumour heterogeneity directly from histology slides,” says Phedias. “It’s a useful tool to complement other molecular approaches and contribute to ongoing personalized medicine efforts.”
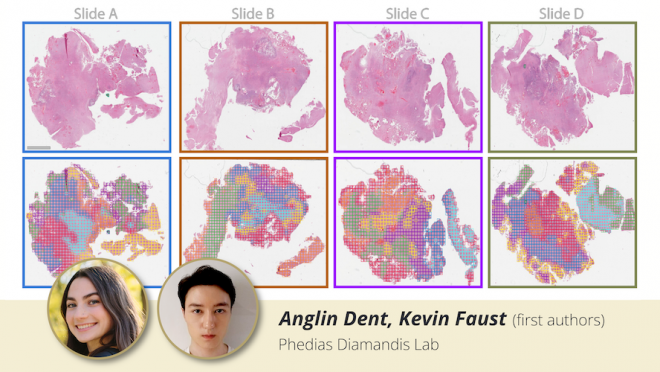

Using deep neural networks to map tumour histomorphology variations. The first row of images are H&E histological slides from a high-grade glioma and the second row includes their respective HAVOC heterogeneity maps. The coloured clusters proposed by HAVOC map are intra-tumour variations in cancer biology that can one day help inform custom therapy design for patients (from Science Advances).
“The danger of AI is that we are not really sure how it behaves in new situations,” says Phedias.
“When humans encounter an unexpected situation, we know it's better to be cautious and avoid a mistake rather than trying to fit it into a mathematical formula, but this is not the case for AI.”
Taking a rare brain tumour as an example, a case that only occurs once every ten years can still fire up the neuronal connections in a pathologist’s brain to alert them to be mindful when making a diagnosis. However, when a computer encounters a rare tumour, it can make a guess based on probability and problems arise. When adding up the probabilities of running into different types of rare tumours, it can mount up to 10-20% of the total cases and it is very difficult to train a system to properly react to them.
“In addition to a large training data set, it's also important to consider the creator's artistic ability to develop intelligent algorithms that perform well in specific circumstances,” says Phedias. “We need to rely on the community through crowdsourcing to solve some of those rare circumstances.”
To better include rare cases and expand HAVOC’s application in different disease areas in pathology, Phedias’ team has developed PHARAOH (PHenotyping And Regional Analysis Of Histology), an open online platform that makes the HAVOC map accessible and easy to generate.
Researchers and pathologists can tweak the parameters to design and develop their own AI tools on PHARAOH, based on their specific pathology expertise and needs. PHARAOH allows users to share their creations with the entire community to avoid duplicating efforts.
“The motivation behind PHARAOH is to enable clinicians to contribute and utilize computational pathology without the need to have a deep understanding of the advanced coding needed to design AI algorithms,” says Phedias.
Phedias explained how to utilize PHARAOH in this YouTube video in detail.
In 2019, Phedias created the YouTube channel NeuroscIQ, an online space to foster education and curiosity within the world of neuroscience. He and motivated students in his lab have been contributing to the channel over the years, sharing their most updated research information, and dissecting neuroscience breakthroughs for lay audiences.
“It has been something I wanted to do since my residency, because I’m fascinated by the idea of disseminating knowledge quickly in a free and scalable manner to anyone who wants to consume it. That is how I learned.”
“It took me eight years to take the action, and I wish I started earlier,” Phedias recalls. “When I was contemplating it, I feared that people might not receive my content well. But it turns out, people generally see sharing as a positive thing and they are appreciative to hear other people’s points of view.”
The complexity of neuroscience initially deterred Phedias from studying the brain systematically, until it became a necessity for his PhD work.
The daunting task of starting a YouTube channel was not realized for eight years until his motivation to share eventually overcame perfectionism and the fear of potential critique.
The rise of AI poses an uncertainty for the future but Phedias jumped in to make it a tool that can empower the field.
Each time when facing unfamiliarity, the fear does not go away, but it takes Phedias less time than before to react to it.
“If you're scared of something, it's probably something that will make you stronger if you eventually embrace it,” says Phedias.
Phedias trained for a marathon once. Eight weeks in, he went from only being able to run two kilometers on the first practice, to finishing the entire 42 kilometers on the day of the marathon. This training experience has shed light for him on how to approach the daunting quests in life.
“I don't get as overwhelmed trying to learn a new skill as I did before,” says Phedias. “During training, you never actually fully run a marathon. You just need to get yourself to a specific level where mentally you know that this can be done and you can complete it.”
“Some of the biggest barriers are not our capabilities, but our mental approach to the problem in front of us. My training partner at the time gave me some good advice. If something seems outside of your current capabilities, start by breaking it into smaller pieces that you know you can achieve. For running, he would say, just try and get to the next lamppost on the road. Once you get there, then just do that again and again until the run is over.”
Meet PMResearch is a story series that features Princess Margaret researchers. It showcases the research of world-class scientists, as well as their passions and interests in career and life—from hobbies and avocations to career trajectories and life philosophies. The researchers that we select are relevant to advocacy/awareness initiatives or have recently received awards or published papers. We are also showcasing the diversity of our staff in keeping with UHN themes and priorities.
Research conducted at UHN's research institutes spans the full spectrum of diseases and disciplines, including cancer, cardiovascular sciences, transplantation, neural and sensory sciences, musculoskeletal health, rehabilitation sciences, and community and population health.
Learn more about our institutes by clicking below: